Dilute Libraries to the Starting Concentration
This step dilutes libraries to the starting concentration for your sequencing system and is the first step in a serial dilution. After diluting to the starting concentration, libraries are ready to be denatured and diluted to the final loading concentration. For details on sequencing system compatibility, refer to the Illumina support site.
The final loading concentrations are a starting point and general guideline. Optimize concentrations for your workflow and quantification method over subsequent sequencing runs or by flow cell titration.
| 1. | Calculate the molarity value of the libraries. Use the following formula: |
Use 200 bp as the average library size. The formula uses 660 g/mol as the average weight for a single DNA bp.
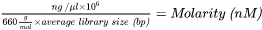
| 2. | Using the molarity value, dilute libraries to the starting concentration for your system. |
If pooling, combine libraries with different sample indexes in equimolar amounts if similar sequencing depth is needed for each library.
|
Sequencing System |
Starting Concentration (nM) |
Final Loading Concentration (pM) |
|---|---|---|
|
iSeq 100* |
4 |
75 |
|
MiSeq* |
4 |
10 |
|
MiniSeq* |
4 |
1.2 |
|
NextSeq 500/550 |
4 |
1.2 |
|
NextSeq 1000/2000 |
2 |
[Onboard Denature & Dilute] 650 [Manual Denature & Dilute] 75 |
|
NovaSeq 6000 |
10 |
200–300 |
* End of life announced. Refer to the Illumina website for more information.
| 3. | Follow the denature and dilute instructions for your system to dilute libraries to the final loading concentration. |
| • | For iSeq 100, refer to the iSeq 100 System Product Documentation. |
| • | For the following sequencing systems, refer to the Denature and Dilute Protocol Generator and follow the Standard loading protocol: |
| • | MiSeq |
| • | MiniSeq |
| • | NextSeq 500/550 |
| • | NovaSeq 6000 |
| • | For NextSeq 1000/2000, refer to the Denature and Dilute Protocol Generator and follow the Onboard or Manual protocol. |
| 4. | Prepare and load the pooled library on the sequencing system according to the specific Illumina sequencing system product documentation. |
| • | Although not required, you can add PhiX to the sequencing run. For the optimal PhiX percentage, refer to the denature and dilute instructions for your system. |
| • | Illumina miRNA Prep libraries require 1 x 72 bp read length with 10 bp dual indexing. Illumina recommends allocating 5–10 million reads per sample. |
