HiSeq/GAIIx v5 Sample Sheet Generation
This documentation has been updated and moved to the Illumina Integration Overview section. See CASAVA 1.8.2 Sample Sheet Generation.
Package Version: BaseSpace Clarity LIMS HiSeq/GAIIx Package v5.1 or later
Compatibility: BaseSpace Clarity LIMS v3.0 or later; NGS Extensions Package v5.0 or later. Recent features are marked with the package version in which they first appear.
New in HiSeq 5.1: -useSampleLimsID (-s) and -appendLimsID (-a) options.
The createSampleSheet script creates sample sheets for Illumina HiSeq and GAII instruments that are compatible with CASAVA 1.8.2.
Typically, sample sheet generation is run on the Cluster Generation step and the script is capable of creating one sample sheet per flow cell loaded during the step.
Installation and Upgrade Notes
To take advantage of the new features BaseSpace Clarity LIMS HiSeq/GAIIx Package v5.1 provides, customers who are installing or upgrading to the 5.1 package must manually modify the Cluster Generation (Illumina SBS) 5.0 process type configuration using the Operations Interface.
New features included in the 5.1 package enable the following:
| • | An option for populating the sample sheet SampleID field with the sample LIMS ID instead of name. |
| • | An option for populating the sample sheet SampleID field with the protocol step LIMS ID appended to the end of the value entered. This option will ensure unique FASTQ file names per sequencing run when these are generated by Illumina. |
To enable sample sheet generation for multiple flow cells:
| • | The Cluster Generation step must be configured to produce one shared result file per sample sheet that will be created (see Support for Multiple Sample Sheets). |
| • | The EPP command line must specify the -c parameter with the LIMSID of the shared result file that will store each sample sheet in BaseSpace Clarity LIMS. The sample sheets will be attached to these result files and named as per the corresponding flow cell. |
To enable unique FASTQ file names per sequencing run:
| • | The Cluster Generation step EPP command line must be configured to use the -useSampleLimsID (-s) and -appendLimsID (-a) options. See Script Parameters and Usage |
| – | useSampleLimsID ensures unique entries in the SampleID column by using the sample LIMS ID instead of its name |
| – | appendLimsID ensures unique names per run by appending the LIMS ID of the current protocol step |
|
Field Name |
Field Type |
Required? |
Notes |
|
Reference Genome |
Text |
No |
Supplies data for sampleRef column in sample sheet. |
The following table defines the parameters used by the createSampleSheet script.
|
-u |
LIMS username (Required) |
|
-p |
LIMS password (Required) |
|
-i |
|
|
-c, -csvFileLimsIds |
Sample sheet CSV file LIMS ID (Required - repeats allowed) |
|
-e, -errorLogFileName |
Log file name (required) |
|
-l, -useProjectLimsID |
Accepted values: true or false. Provide with quotes e.g. -l 'true' (Optional) Project LIMS ID will be used instead of the project name in the sample sheet (Optional) (lower case L) |
|
-s, -useSampleLimsID |
Accepted values: true or false . Provide with quotes e.g. -s 'true' (Optional) Sample LIMS IDs will be used instead of Sample Names in the sample sheet (HiSeq/GAIIx 5.1) |
|
- a, -appendLimsID |
Accepted values: true or false . Provide with quotes e.g. -a 'true' (Optional) The LIMS ID of the protocol step will be appended to the Sample Name values in the sample sheet. Use this option to guarantee unique FASTQ file names per run. (HiSeq/GAIIx 5.1) |
Below is the default command line that ships with the Cluster Generation (Illumina SBS) 5.0 step in new installations. The createSampleSheet portion of the parameter string is shown in bold.
bash -c "/usr/bin/java -jar /opt/gls/clarity/extensions/Illumina_HiSeq/v5/EPP/hiseqgaii-extensions.jar -u {username} -p {password} -i {processURI:v2:http} script:setUDF -f 'Progress' -t '//input/@uri->//sample/@uri' -v 'Library ready for sequencing' script:createSampleSheet -c {compoundOutputFileLuid1} -c {compoundOutputFileLuid2} -c {compoundOutputFileLuid3} -c {compoundOutputFileLuid4} -c {compoundOutputFileLuid5} -e {compoundOutputFileLuid6} script:labelNonLabeledOutputs -l 'NoIndex' script:initArtifactUDFs"
Any one-dimensional container types with both numeric rows and numeric columns are supported.
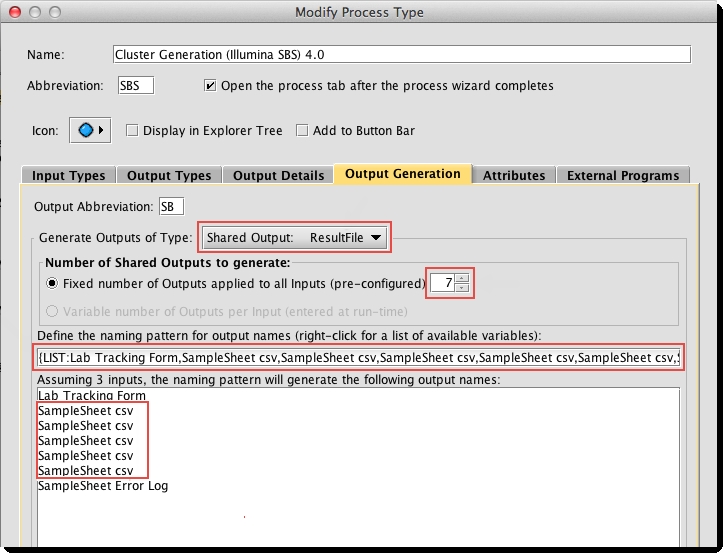
Support for Multiple Sample Sheets
If generating multiple sample sheets, you must update the process to create placeholders for each of the sample sheets, as illustrated below.
|
Info Messages |
Reason/Condition |
|
[Info] – Added data line for input: {input name} |
For each line of data added to the sample sheet. |
|
[Info] – Successfully generated Illumina HiSeq Sample Sheet. |
On successful completion with no warnings or errors. |
If any of the errors described in the following table are encountered:
| • | The script will terminate. |
| • | An error log file will be written and attached to the step |
| • | No sample sheet will be generated. |
|
Error Messages |
Reason/Condition |
|
Unable to determine {user or password} |
Unable to determine FTP credentials for ftp.user or ftp.password. |
|
There are {number of flow cells created by the step} Flow Cell(s), but this step only supports sample sheet creation for {number of -c parameters pass on the command line} Flow Cell(s). Please restart the step with fewer samples. |
More flow cells were found in the step than result file outputs in which to save the sample sheets. This can be remedied by changing the step configuration to accommodate additional files or by running the step with fewer inputs. |
|
Container type not supported {containerType name} for input {input name}. The expected container type must be a single column or single row Flow Cell. |
Unsupported container type used as a flow cell. Will occur if container has two dimensions. |
|
Well coord not supported {input node xml} for input {input name}. The expected format is [numeric]:[numeric]. |
Unsupported well format for container type used as a flow cell. |
|
Invalid number format for {placement string} for input {input name}. The expected format is [numeric]:[numeric]. |
Unexpected error caught trying to parse the well coordinate. Ensure the flow cell has numeric wells and numeric columns. |
|
Unable to find reagent type {reagent label on input} for input {input name}. |
There is no reagent type configured with the name of the reagent label found on the artifact. |
|
Unable to find Index special type with Sequence attribute for input {input name}. Please check Reagent Type configuration of {reagent label on input}. |
There is no reagent type configured with a special type of Index that has a Sequence attribute. |
| • | The contents of the sample sheet are ordered by Lane, then by Description (LIMS ID). |
| • | Project and sample names in the sample sheet cannot contain illegal characters that are not allowed by some file systems. These characters are the space character and the following: ? ( ) [ ] / \ = + < > : ; " ' , * ^ | & |
| • | The script will replace these characters with an underscore "_". |
|
Column Header |
Description |
|
FCID |
The name of the flow cell container. |
|
Lane |
The flow cell lane number corresponding to the input placement. |
|
SampleID |
Information from the submitted sample of the input, or the submitted sample of the (first) upstream pooling input if pooling exists. Will be sample name or LIMS ID, depending on command-line value. An additional command line option allows the step LIMS ID to be appended to the end of this value (e.g., "Sample1-1234"). See Script parameters and usage. |
|
SampleRef |
The value of the Reference Genome UDF on the input's sample. |
|
Index |
Uses the “Sequence” attribute value from index reagents. |
|
Description |
The LIMS ID of the step input, or the LIMS ID of the upstream pooling input if pooling exists. |
|
Control |
An input is considered a control if its name contains PhiX (case-sensitive). |
|
Recipe |
This column is always blank. |
|
Operator |
The name of the technician who initiated the step in the LIMS. |
|
SampleProject |
The project name of the Illumina Sequencing Analysis process input, or the project name of the upstream pooling input if pooling exists. If the '-l' parameter is true, the project LIMS ID will be used instead of the project name. |
