Homologous Recombination Deficiency
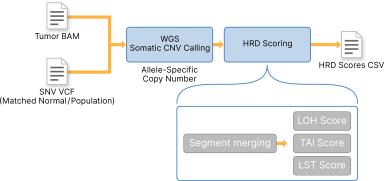
DRAGEN Homologous Recombination Deficiency (HRD) Scoring takes in allele-specific copy number calls in either VCF format or directly streamed from somatic copy number callers. DRAGEN HRD then calculates scores for Loss of Heterozygosity (LOH), Telomeric Allelic Imbalance (TAI), and Large-Scale State Transition (LST). The three scores are output to the .hrdscore.csv file. You can only use DRAGEN HRD when inputting results from WGS somatic CNV calling.
Use the following command-line options to run HRD scoring. You can run HRD scoring with somatic CNV calling or after using somatic CNV calling results.
To run HRD scoring together with somatic CNV calling, use the following options. See Somatic CNV Calling for more parameters.
| • | --enable-hrd—Set to true to enable HRD scoring to quantify genomic instability. |
| • | --enable-cnv—Set to true to enable CNV calling to run together with HRD scoring. |
To run HRD scoring after somatic CNV calling, use the following options:
| • | --enable-hrd—Set to true to enable HRD scoring to quantify genomic instability. |
| • | --hrd-input-ascn —If running HRD scoring after somatic CNV calling, specify the allele-specific copy number file (*cnv.vcf.gz). |
| • | --hrd-input-tn—If running HRD scoring after somatic CNV calling, specify the tumor normalized bin count file (*.tn.tsv.gz). |
The following metrics are included in the .hrdscore.csv output file. The following is an example output file.
|
Sample |
LOH_Score |
TAI_Score |
LST_Score |
HRD_Score |
|---|---|---|---|---|
|
Sample |
16 |
17 |
28 |
61 |
